Modern Research Techniques of Apoptotic Cell Death (Review)
The review concerns modern research techniques of apoptotic cell death in vitro and in vivo. Apoptosis monitoring has been shown to be based on recording characteristic cell changes: integrity loss and depolarization of plasma membrane, phosphatidylserine and phosphatidylethanolamine exposure on cell membrane surface, activation of caspases, mitochondrial membrane depolarization, DNA fragmentation, the changed level of biochemical markers. The major techniques for in vitro apoptosis studies are light and electron microscopy, flow cytometry, fluorescent microscopy, immunohistochemistry, enzyme immunoassay, Western blot, electrophoresis and TUNEL technique. Currently, noninvasive techniques used to study apoptosis in vivo are of great importance since they are capable of recording the process in tissues, organs and the whole body: positron emission tomography (PET), single-photon emission computed tomography (SPECT), magnetic resonance tomography (MRT), fluorescent spectroscopy, FLIM/FRET-imaging. Great attention is paid to the search for contrast agents specific to various molecules of an apoptotic cascade. The nature of apoptotic reactions has been demonstrated to serve as a marker of chemotherapeutic efficiency and help choosing chemotherapy. A combination of several techniques to study apoptotic cell death has been found to gain its popularity, e.g. SPECT and optical imaging, PET and optical imaging, PET and MRT. In the long term, by means of these techniques it will be possible both to determine the disease severity and also assess the treatment efficiency.
Apoptosis is a process of a programmed cell death, which is controlled genetically and typical for all mammalian cells [1–3]. It plays a key role in cell growth and differentiation, the development of the whole body and tissue homeostasis. The knowledge of molecular mechanisms participating in apoptosis initiation and execution contributes to the understanding of the processes involved in the development of many diseases including oncological ones [3, 4]. Currently, the cell death regulatory factors are considered as the primary targets of antitumor action, and a type of apoptotic reactions can serve as a marker of rational chemotherapeutic efficiency and help choosing chemotherapy [4–7].
Now, a great number of methods enabling to discover many mechanisms of apoptosis have been suggested for qualitative and quantitative analysis of apoptotic reactions in a cell [8]. These techniques are based on recording typical cell alterations: morphological, biochemical and molecular (Figure 1) [8, 9]. There has been gained a great deal of evidence on apoptotic cell death mechanisms; however, currently, most studies concerned with cell death have been performed on cell cultures in vitro. In recent years, there is an active search and development of noninvasive techniques to investigate apoptosis in vivo suitable for studing this process in tissues, organs and in a whole body [8, 10].
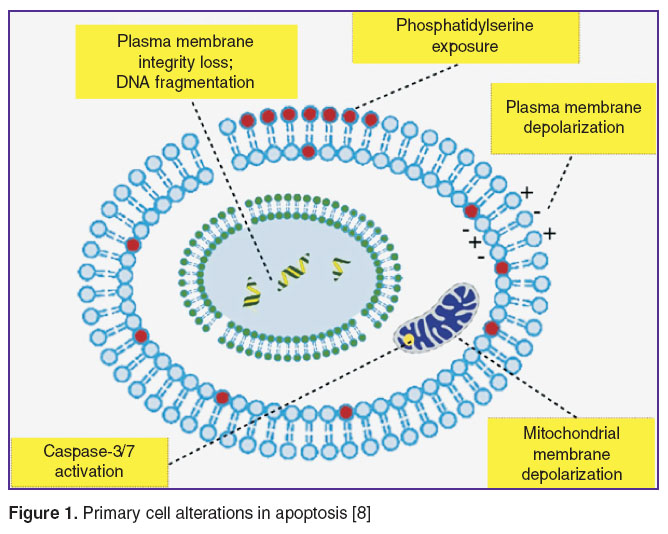 Figure 1. Primary cell alterations in apoptosis [8] Figure 1. Primary cell alterations in apoptosis [8]
|
Morphological changes and plasma membrane depolarization
Morphologically, apoptosis involves reduced cell volume, plasma membrane shrinkage, chromatin condensation, nuclear DNA strand breaks followed by nuclear decomposition, cell fragmentation into membrane vesicles with intracellular content (apoptotic bodies) phagocytized by macrophages and adjacent cells [5].
The most available and easiest method to detect apoptotic cells and study their morphological characteristics is light microscopy of histological specimens. For this purpose thin-layer tissue sections are used stained by Azure A or hematoxylin and eosin [11–14].
Ultrastructural changes typical for apoptotic cells can be detected by electron microscopy. Ultrastructural peculiarities of cells in apoptosis activated by various inductors have been demonstrated by this method. Electron microscopy is generally considered to be more reliable technique to study apoptosis compared to light microscopy [15, 16].
Morphological changes can also be studied by flow cytometry. The reduction of light scattering intensity has been found at early apoptotic stages due to cytoplasm shrinkage. However, changes of light scattering are not specific markers in the development of any cell death pathways; therefore, this analysis is to be combined with other methods providing precise identification of a molecular profile of cell death [17].
The capabilities of positron-emission tomography (PET), single-photon emission computed tomography (SPECT) have been demonstrated to assess the damaged structure and depolarization of plasma membrane in apoptosis in vivo [10, 18]. The approach is based on the use of radio-labeled substances specific to different molecules and accumulated in apoptotic cells. In recent years, a great number of radioactive isotopes using various chelators and co-ligands have been developed. In particular, to study depolarization of cytoplasmic membrane and intracellular pH in apoptotic cell death, a family of small molecules ApoSense has been developed [10, 18, 19]. In 2008, a radioactive nuclide 18F-ML-10 (18F-labeled 5-fluoropentyl-2-methylmalonic acid) was developed on the basis of these molecules to estimate cell membrane depolarization by PET [20, 21]. The first human test showed its safety, stability and fast elimination [22, 23]. Currently, clinical trials using 18F-ML-10 are being carried out to study plasma membrane depolarization in apoptosis of tumor cells and assess chemotherapy efficiency [24].
To image apoptosis by SPECT in vivo, a radioactive nuclide 123I-labeled ML-10 was suggested. There has been shown its absorption by Daudi tumor cells depending on pH level in mice when treated by cyclophosphamide and radiotherapy [25].
Phosphatidylserine and phosphatidylethanolamine exposure on plasma membrane surface
Phospholipids of plasma membrane are characterized by asymmetric distribution: phosphatidylcholine and sphingomyelin are located on an outer side of a lipid bilayer, while phosphatidylserine (PS) and phosphatidylethanolamine (PE) are located on its inner surface. In apoptosis the asymmetry breaks, PS and PE appear on a cell membrane surface and can serve as an indicator of apoptosis initiation [7, 8, 18, 26].
A compound used as a marker of apoptotic cells is recombinant protein, Annexin V, with high affinity for PS. Annexin V is a Ca2+-dependent protein with molecular weight of 35–36 kDa [18, 27]. Flow cytometry and fluorescence microscopy using Annexin V conjugated with a fluorescent dye, e.g., FITC, are applied to detect PS [10, 28]. However, due to the fact that PS displacement from an inner surface of cytoplasmic membrane to the outer one occurs in necrosis as well, Annexin V has been used with such cationic dyes as 7-ADD or propidium iodide (PI), which bind to intact DNA, but they are able to penetrate inside a cell only if cytoplasmic membrane integrity is damaged. Combined staining by Annexin V-FITC and PI enables to identify living cells (Annexin V–/PI–), early proapoptotic changes (Annexin V+/PI–), late apoptosis stage accompanied by secondary necrosis of cells, and a necrotic type of cell death (PI+). Thus, the technique enables to distinguish living cells from those in apoptosis, and from necrotic cells [17, 28, 29].
Due to high affinity of Annexin V for membrane PS, there have been developed various radioactive isotopes with Annexin V to image PS by PET and SPECT in vivo [9, 10]. The most extensively studied and widely used radioisotope for apoptosis detection is 99mTc-labeled-Annexin V, which has found its application in oncology [18, 30]. Currently, 99mTc-HYNIC-Annexin V is the only clinically tested radioisotope: it has undergone phases 2 and 3 of clinical trials in the treatment of patients with lung cancer [31, 32]. All patients were found to have increased uptake of Annexin V by a tumor; moreover, there were significant correlations between Annexin V uptake and treatment results. 18F- and 68Ga-labeled Annexin V were also developed to study apoptotic cell death in vivo [10, 18, 33] (Figure 2). Nevertheless, the use of Annexin V in vivo presents such difficulties as slow pharmacokinetics and high immunogenicity.
 Figure 2. Apoptosis imaging by PET using 18F-labeled Annexin V in mice with squamous cell carcinoma of the head and neck UM-SCC-22B before and after doxorubicin treatment [33] Figure 2. Apoptosis imaging by PET using 18F-labeled Annexin V in mice with squamous cell carcinoma of the head and neck UM-SCC-22B before and after doxorubicin treatment [33]
|
In recent years, there has been the search for low-molecular compounds for apoptosis analysis [34]. In 1995, PSBP-0, a peptide consisting of 14 amino acids, was derived from PS carboxylase [35]. In 2011 it was modified and acquired high affinity for PS [36]. The studies have shown the level of a radioisotope SAAC(99mTc)-PSBP-6 in mice with melanoma to be higher after therapy. It indicates that the peptide can be used to study an early tumor response to chemotherapy [37].
Duramycin, a tetracyclic peptide consisting of 19 amino acids, with molecular weight of 2 kDa, was developed to identify PS on cell membrane surface by PET [38]. In 2008 99mTc-labeled Duramycin with HYNIC as a ligand was used to image apoptosis in vivo [39, 40]. Later on, 18F-labeled Duramycin — [18F]FPDuramycin — was suggested as a novel radioisotope for apoptosis imaging [41]. It was successfully used to study the apoptosis of tumor cells after cyclophosphamide and cisplatin therapy in mice by PET. However, pharmacokinetics showed accumulation of [18F]FPDuramycin in the liver and spleen that limits its application in vivo.
Another technique used to study apoptosis in cells in vivo is optical imaging [42, 43]. Recently developed optical imaging methods provide apoptosis imaging in animal tissues and a whole body [10]. In recent years, increasingly greater attention of researchers in this field is given to the combination of different near infrared fluorophores and ligands [44]. Annexin V labeled by Cy 5.5, a near infrared fluorophore, was used for apoptosis optical imaging in vivo [45]. In particular, it was used to study apoptosis in tumors during chemotherapy by fluorescence tomography. The increase of a fluorescent signal was found in treated tumors compared to the control [46].
Magnetic resonance imaging (MRI) is also applied for apoptosis imaging in vivo. Biotinylated Annexin V bound to Gd-DTPA-labeled liposomes is used to identify PS on plasma membrane surface [47, 48]. Supermagnetic iron oxide nanoparticles conjugated with C2A-domain of Synaptotagmin, protein specific to PS, are used for MRI [49]. Apoptosis imaged by MRI was demonstrated in isolated tumor cells in vitro and in the tumors in vivo treated by chemotherapeutic agents. Higher contrast accumulation has been shown in a tumor compared to the control [50].
Caspase activation
In most cases, in animal and human cells apoptosis is related to proteolytic activation of a caspase cascade. Caspases is a family of evolutionarily conserved cysteine proteases, which specifically cleave a peptide bond formed after aspartic acid residues [51]. Caspases are divided into two main groups: caspases-1 (caspase-1, 4, 5, 13 and 14) and caspase-2 playing a key role in the apoptosis process (caspase-2, 3, 6–10). The second group is subdivided into initiator (caspase-2, 8, 9 and 10) and effector (caspase-3, 6 and 7) caspases [51, 52]. Different mechanisms of apoptosis induction through membrane receptors or mitochondria cross at caspase-3 level, therefore the measurement of its activity is the most informative method to reveal apoptosis [6, 10].
Caspase-3 activity can be determined by immunohistochemistry (IHC), enzyme-linked immunosorbent assay (ELISA) or Western blotting using mono- and polyclonal antibodies specific to caspase-3. However, these techniques do not make it possible to record the enzyme activation changes [28, 29, 53–55].
Flow cytometry using high-specific peptide substrates or inhibitors labeled by fluorophore is a widely applied technique to study caspase activity [17].
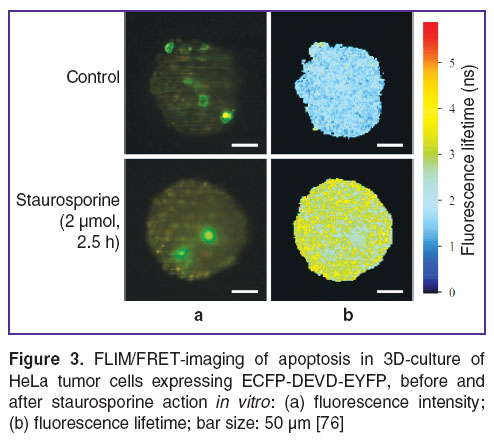
In recent years, the changes of apoptosis in living cells are imaged by fluorescence imaging (microscopy, tomography) and fluorescence spectroscopy. For this purpose fluorescent proteins are widely used as markers of caspase-3 activation. The main approach is the usage of the so called FRET (Förster resonance energy transfer) sensors consisting of a pair of proteins (a donor and an acceptor) bound by a peptide sequence DEVD. In bound state, an exited donor transfers energy by a resonance mechanism to an acceptor resulting in a donor’s reduced fluorescence intensity, while acceptor's fluorescence increases [56–63]. In addition, fluorescence lifetime of a donor in FRET-pair is lower compared to its free state [59, 64]. When apoptosis and caspase-3 activation are initiated, DEVD sequence is cleaved, and FRET-reaction becomes impossible. The most popular FRET-sensor to detect caspase-3 activation is SCAT3 consisting of proteins ECFP and Venus [63, 65–69]. Recently, there has been suggested a sensor to study simultaneously the activity of caspase-8 and caspase-3 consisting of fluorescent proteins seCFP, Venus and mRFP1 bound by the linkers IETD and DEVD [70]. The use of FRET-sensors and fluorescence imaging techniques enable to observe spatiotemporal dynamics of caspase-3 activation in individual living cells in vitro and in vivo by the change of fluorescence intensity or lifetime. It should be noted that time-resolved fluorescence imaging FLIM (fluorescence lifetime imaging microscopy) is considered to be one of the most precise methods to assess FRET-reaction efficiency [59, 66, 71–77] (Figure 3). In particular, apoptosis activation using chemotherapeutic agents in different tumor cells in vitro and in vivo has been shown by this technique [58, 59, 66].
Light penetration depth is of primary importance for in vivo optical imaging of apoptosis in organs and tissues. Animal tissues are known to be more transparent in red and near infrared spectral range, therefore, in recent years, researchers have aimed their efforts at developing fluorescent proteins and FRET-sensors working in this range. The following fluorescent proteins emitting in an orange-red range have been isolated: mRFP, Katushka, mKate2, IFP1.4, iRFP670, iRFP682, iRFP702, iRFP720, etc. [78–80]. Their usage enables to increase imaging depth and reduce the effect of autofluorescence of tissues and a toxic effect of probing radiation [81–86].
Another approach to the assessment of caspase activity consists in the usage of different radioisotope-labeled substances. In 2001 there were appeared new data on activity and selectivity of some isatin analogs of caspase-3. In 2008, 18F-labeled analog of isatin was synthesized and described. A radioisotope [18F]ICMT-11 was used to study apoptosis in vitro and on animals in vivo by PET. In vivo study using [18F]ICMT-11 revealed higher caspase-3 activity in tumor cells after cyclophosphamide therapy. Moreover, a signal was increasing within the first 24 h after chemotherapy in mice with B-cell lymphoma 38C13 [87]. Preclinical studies of [18F]ICMT-11 showed its safety for human [88].
Progress in magnetic resonance imaging 1H-MRI is being carried out to image the activity of apoptotic cascade enzymes. However, a significant limitation of the technique is a high background signal, therefore, many researchers pay much attention to the use of 19F-MRI. 19F-MRI-contrast Gd-DOTA-DEVD-Tfb consisting of Gd3+-complex, a peptide group DEVD and 19F-containing group (Tfb) was synthesized to determine caspase-3 activity. When DEVD-sequence is cleaved by caspase-3, 19F-MRI-peak is higher and sharper indicating the disturbance of a paramagnetic effect between Gd3+ and 19F [10].
Mitochondrial membrane depolarization
Mitochondria are known to play a key role in apoptosis [10, 89, 90]. The damage of mitochondrial membrane potential Δψm and permeability of mitochondrial membrane can result from an oxidative stress or chemotherapy leading to the release of membrane-associated proteins including cytochrome c [91, 92]. Mitochondrial membrane depolarization occurs at early apoptotic stages and before the primary morphological and biochemical alterations. Membrane potential monitoring can give the information about the kinetics of apoptotic processes in the tissues under study [8].
Membrane-bound lipophilic cationic fluorophores, in particular rhodamine 123 (Rh123); 3.3’-dihexyloxacarbocyanine (DiOC6); 5.5’,6.6’-tetra-chloro-1.1’; 3.3’-tetraethylbenzimidazole-carbocyanine iodide (JC-1) and other fluorescent dyes (e.g., TMRE, TMRM) are used to detect Δψm due to their capability to accumulate in mitochondria as probe molecules; they enable to record Δψm by fluorescence intensity [28, 29, 55, 93]. Moreover, Δψm analysis is carried out both in single cells and also in isolated mitochondria by flow cytometry.
In 2007, 18F-labeled phosphonium cation (18F-FBnTP) was used for noninvasive study of mitochondrial membrane potential by PET [94]. Phosphonium cations can penetrate through a lipid bilayer due to their lipophilicity and a positive charge, and accumulate inside mitochondria. When membrane potential is lost during apoptosis, the concentration of phosphonium cations in a cell decreases. In 2009, 18F-FBnTP was applied in treatment of prostate carcinoma in mice in vivo [95].
DNA fragmentation
During apoptosis, under the action of endonucleases, there are numerous DNA strand breaks resulting in the formation of many 3’-ends [96]. TUNEL technique (terminal deoxynucleotidyl transferase-mediated dUTP-biotin nick-end labeling), or terminal deoxynucleotidyl end labeling is widely applied to reveal the process [55]. The principle of TUNEL technique is in specific binding to 3’-end of a broken biotin-labeled deoxyuridine triphosphate DNA strand. Such binding is catalyzed by deoxynucleotidyl transferase. The method is considered to be safe and specific for apoptosis detection, since it aims at verification of a primary event: DNA breakdown under Mg2+/Ca2+-dependent endonucleases resulting in the formation of fragments, their size being multiple of a nucleosome [10–12, 97, 98]. One of recent modifications of TUNEL technique enables to detect DNA fragmentation in a sample containing just 5 ng of DNA, so that the sensitivity of the method is over 200 times as high than a routine staining procedure.
It should be noted that oligonucleosome degradation of chromatin is revealed well before apoptosis signs become visible by means of light microscopy, and can be detected by electrophoresis [99, 100]. A discrete “ladder”, the fragments of which have the size multiple of 200 base pairs (one nucleosome), can serve as an evidence of apoptosis that can be proved by markers of molecular weight. The necessity of such control arises from the fact that when DNA is damaged by deoxyribonucleases observed in necrosis, “a ladder” can also be revealed though the ladder is oligonucleosome one rather than oligonucleotide, the weight of the fragments being multiple of one base. These conditions necessitate controlling the results of TUNEL and DNA electrophoresis by light microscopy with the verification of apoptotic morphological signs.
The release of mono- and oligonucleotide DNA fragments from the nucleus in cytoplasm of apoptotic cells can be studied by IHC and ELISA using mono- and polyclonal antibodies specific for nucleosome DNA fragments contained in lysate of apoptotic cells [100]. The sensitivity of the method is over 500 times as high compared to DNA fragmentation revealed by gel-electrophoresis. An essential advantage of the technique is the possibility to analyze simultaneously more samples, while the main disadvantage is the necessity to analyze fresh samples, since their storage can result in significant weakening of reactivity.
Frequently, to detect apoptotic cells, fluorescence microscopy with fluorescent dyes is used, the dyes being specifically bound to DNA: DAPI, Hoechst 33342, acridine orange and ethydium bromide [63, 68]. Moreover, both vitally stained cells in suspension and fixed preparations are studied. The detection of condensed chromatin fluorescing brightly is a usual apoptotic sign in fluorescence microscopy. Apoptosis in cell suspension can be measured quantitatively by combining DNA treatment with specific dyes and flow cytometry. Propidium iodide is frequently used for the treatment [16, 17].
SPECT has been shown to be used to detect DNA damage in apoptosis in vivo [101]. Damage is detected by a signal protein gH2AX, which binds to a damaged DNA strand in apoptosis, and can be found by fluorophore- or 111In-DOTA-labeled antibodies. The signal has been shown to increase in SPECT after bleomycin therapy of breast cancer on animal models.
Biochemical markers of apoptosis
IHC, ELISA and Western blotting are applied to identify proteins composing a cascade of biochemical processes leading to apoptosis [54]. Based on these considerations a conclusion is made about cell possibility to come into apoptosis but not about the apoptosis level in a cell population [16, 29, 67, 102]. Poly- and monoclonal antibodies to proteins p53 — DO-7, РАВ-1801 (to detect wt p53 and mt p53), РАВ-240 (to identify mt p53 alone), etc. are normally used. In addition to antibodies to p53, immunoglobulins to other key points of the genetic program of apoptosis are used: bcl-2, bax, MPM-2, RB, Fas (monoclonal antibodies ICO-160, anti-Fas, anti-APO-1), cyclins, caspases, etc.
Fluorescence imaging and FLIM using FRET-sensors to proapoptotic factors are used for apoptosis imaging in living cells. In particular, there has been developed a sensor consisting of fluorescent proteins YFP and CFP to determine Bid activation and apoptosis induction [103].
The Table represents the information about different techniques to detect apoptotic cell death in vitro and in vivo.
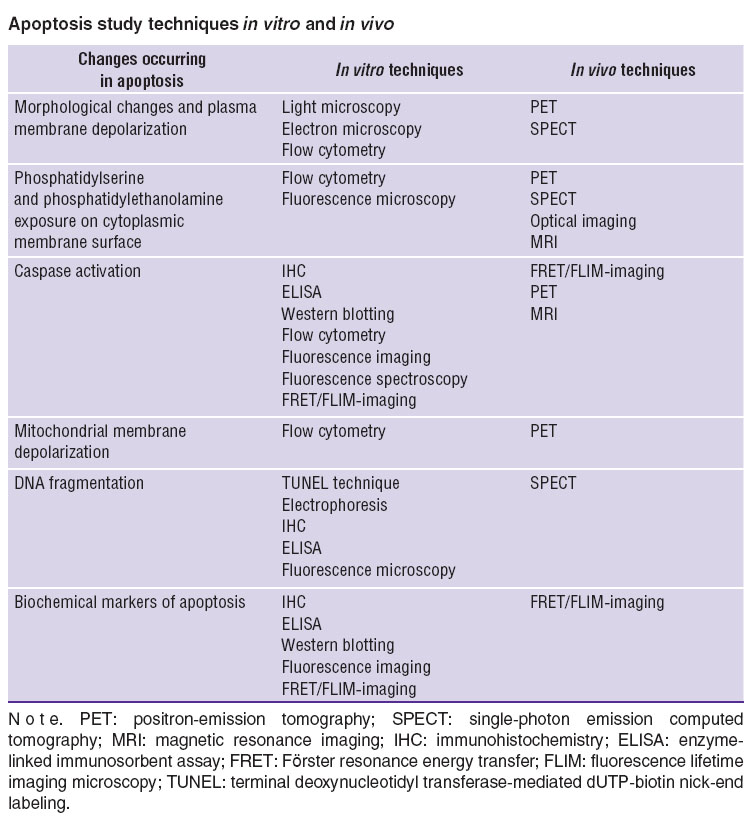 Apoptosis study techniques in vitro and in vivo Apoptosis study techniques in vitro and in vivo
|
It should be noted that recent research in this field deals with the development of apoptosis study techniques in vivo that is of great importance for clinical medicine. A combination of several techniques gains its popularity, e.g., SPECT and CT; SPECT and optical imaging; PET and optical imaging; PET, SPECT and CT; PET and MRI [10, 104–106]. In the long term, by means of such combined techniques it will be possible both to determine the disease severity and detect tumors, and also assess the treatment efficiency and predict recurrences.
Study Funding. The study was supported by Russian Science Foundation (project No.14-25-00129).
Conflicts of Interest. The authors have no conflicts of interest to declare.
References
- Goldar S., Khaniani M.S., Derakhshan S.M., Baradaran B. Molecular mechanisms of apoptosis and roles in cancer development and treatment. Asian Pac J Cancer Prev 2015; 16(6): 2129–2144.
- Labi V., Erlacher M. How cell death shapes cancer. Cell Death Dis 2015; 6: e1675, http://dx.doi.org/10.1038/cddis.2015.20.
- Wong R.S. Apoptosis in cancer: from pathogenesis to treatment. J Exp Clin Cancer Res 2011; 30: 87, http://dx.doi.org/10.1186/1756-9966-30-87.
- Su Z., Yang Z., Xu Y., Chen Y., Yu Q. Apoptosis, autophagy, necroptosis, and cancer metastasis. Mol Cancer 2015; 14(1): 48, http://dx.doi.org/10.1186/s12943-015-0321-5.
- Lopez J., Tait S.W. Mitochondrial apoptosis: killing cancer using the enemy within. Br J Cancer 2015; 112(6): 957–962, http://dx.doi.org/10.1038/bjc.2015.85.
- Huang Q., Li F., Liu X., Li W., Shi W., Liu F.F., O’Sullivan B., He Z., Peng Y., Tan A.C., Zhou L., Shen J., Han G., Wang X.J., Thorburn J., Thorburn A., Jimeno A., Raben D., Bedford J.S., Li C.Y. Caspase 3-mediated stimulation of tumor cell repopulation during cancer radiotherapy. Nat Med 2011 Jul; 17(7): 860–866, http://dx.doi.org/10.1038/nm.2385.
- Smith B.A., Smith B.D. Biomarkers and molecular probes for cell death imaging and targeted therapeutics. Bioconjug Chem 2012; 23(10): 1989–2006, http://dx.doi.org/10.1021/bc3003309.
- Neves A.A., Brindle K.M. Imaging cell death. J Nucl Med 2014; 55(1): 1–4, http://dx.doi.org/10.2967/jnumed.112.114264.
- Bernsen M.R., Kooiman K., Segbers M., van Leeuwen F.W., de Jong M. Biomarkers in preclinical cancer imaging. Eur J Nucl Med Mol Imaging 2015; 42(4): 579–596, http://dx.doi.org/10.1007/s00259-014-2980-7.
- Zeng W., Wang X., Xu P., Liu G., Eden H.S., Chen X. Molecular imaging of apoptosis: from micro to macro. Theranostics 2015; 5(6): 559–582, http://dx.doi.org/10.7150/thno.11548.
- Kasagi N., Gomyo Y., Shirai H., Tsujitani S., Ito H. Apoptotic cell death in human gastric carcinoma: analysis by terminal deoxynucleotidyl transferase-mediated dUTP-biotin nick end labeling. Jpn J Cancer Res 1994; 85(9): 939–945, http://dx.doi.org/10.1111/j.1349-7006.1994.tb02972.x.
- Jolly P.D., Smith P.R., Heath D.A., Hudson N.L., Lun S., Still L.A., Watts C.H., McNatty K.P. Morphological evidence of apoptosis and the prevalence of apoptotic versus mitotic cells in the membrana granulosa of ovarian follicles during spontaneous and induced atresia in ewes. Biol Reprod 1997; 56(4): 837–846, http://dx.doi.org/10.1095/biolreprod56.4.837.
- Drachenberg C.B., Ioffe O.B., Papadimitriou J.C. Progressive increase of apoptosis in prostatic intraepithelial neoplasia and carcinoma: comparison between in situ end-labeling of fragmented DNA and detection by routine hematoxylin-eosin staining. Arch Pathol Lab Med 1997; 121(1): 54–58.
- Chyle V., Pollack A., Czerniak B., Stephens L.C., Zagars G.K., Terry N.H., Meyn R.E. Apoptosis and downstaging after preoperative radiotherapy for muscle-invasive bladder cancer. Int J Radiat Oncol Biol Phys 1996; 35(2): 281–287, http://dx.doi.org/10.1016/0360-3016(96)00089-2.
- Huang R.F., Wei Y.J., Inbaraj B.S., Chen B.H. Inhibition of colon cancer cell growth by nanoemulsion carrying gold nanoparticles and lycopene. Int J Nanomedicine 2015; 10: 2823–2846, http://dx.doi.org/10.2147/IJN.S79107.
- Li Q., Ren F.Q., Yang C.L., Zhou L.M., Liu Y.Y., Xiao J., Zhu L., Wang Z.G. Anti-proliferation effects of isorhamnetin on lung cancer cells in vitro and in vivo. Asian Pac J Cancer Prev 2015; 16(7): 3035–3042.
- Voitkova V.V. Study of apoptosis with use of flow cytometry. Bulletin of the East Siberian Scientific Center SB RAMS 2010; 76(6 Part 1): 220–225.
- Ogawa K., Aoki M. Radiolabeled apoptosis imaging agents for early detection of response to therapy. Scientific World Journal 2014; 2014: 732603, http://dx.doi.org/10.1155/2014/732603.
- Damianovich M., Ziv I., Heyman S.N., Rosen S., Shina A., Kidron D., Aloya T., Grimberg H., Levin G., Reshef A., Bentolila A., Cohen A., Shirvan A. ApoSense: a novel technology for functional molecular imaging of cell death in models of acute renal tubular necrosis. Eur J Nucl Med Mol Imaging 2006; 33(3): 281–291, http://dx.doi.org/10.1007/s00259-005-1905-x.
- Reshef A., Shirvan A., Akselrod-Ballin A., Wall A., Ziv I. Small-molecule biomarkers for clinical PET imaging of apoptosis. J Nucl Med 2010; 51(6): 837–840, http://dx.doi.org/10.2967/jnumed.109.063917.
- Reshef A., Shirvan A., Waterhouse R. N., Grimberg H., Levin G., Cohen A., Ulysse L.G., Friedman G., Antoni G., Ziv I. Molecular imaging of neurovascular cell death in experimental cerebral stroke by PET. J Nucl Med 2008; 49(9): 1520–1528, http://dx.doi.org/10.2967/jnumed.107.043919.
- Höglund J., Shirvan A., Antoni G., Gustavsson S.A., Langström B., Ringheim A., Sörensen J., Ben-Ami M., Ziv I. 18F-ML-10, a PET tracer for apoptosis: first human study. J Nucl Med 2011; 52(5): 720–725, http://dx.doi.org/10.2967/jnumed.110.081786.
- Shirvan A., Reshef A., Allen A., Fenig E., Stenmetz A., Groshar D., et al. Apoptosis imaging with PET-18F-ML-10 for early assessment of response of brain metastases to radiotherapy. J Nucl Med 2009; 50(Suppl 2): S453.
- Allen A.M., Ben-Ami M., Reshef A., Steinmetz A., Kundel Y., Inbar E., Djaldetti R., Davidson T., Fenig E., Ziv I. Assessment of response of brain metastases to radiotherapy by PET imaging of apoptosis with 18F-ML-10. Eur J Nucl Med Mol Imaging 2012; 39(9): 1400–1408, http://dx.doi.org/10.1007/s00259-012-2150-8.
- Bauwens M., de Saint-Hubert M., Cleynhens J., Vandeputte C., Li J., Devos E., Hendrickx S., Ni Y., Reutelingsperger C., Mortelmans L., Mottaghy F.M., Verbruggen A. In vitro and in vivo comparison of 18F and 123I-labeled ML10 with 68Ga-Cys2-AnxA5 for molecular imaging of apoptosis. Q J Nucl Med Mol Imaging 2013; 57(2): 187–200.
- Alam I.S., Neves A.A., Witney T.H., Boren J., Brindle K.M. Comparison of the C2A domain of synaptotagmin-I and annexin-V as probes for detecting cell death. Bioconjug Chem 2010; 21: 884–891, http://dx.doi.org/10.1021/bc9004415.
- Gerke V., Moss S.E. Annexins: from structure to function. Physiol Rev 2002; 82(2): 331–371.
- Lu Y.Y., Chen T.S., Qu J.L., Pan W.L., Sun L., Wei X.B. Dihydroartemisinin (DHA) induces caspase-3-dependent apoptosis in human lung adenocarcinoma ASTC-a-1 cells. J Biomed Sci 2009; 16: 16, http://dx.doi.org/10.1186/1423-0127-16-16.
- Fomin A.S., Koval O.A., Semenov D.V., Potapenko M.O., Kuligina E.V., Kit Yu.Ya. Richter V.A. The analysis of biomedical markers of MCF-7 cells apoptosis induced by recombinant analog of lactaptin. Russian Journal of Bioorganic Chemistry 2012; 38(1): 77–82, http://dx.doi.org/10.1134/s1068162012010086.
- Blankenberg F.G., Katsikis P.D., Tait J.F., Davis R.E., Naumovski L., Ohtsuki K., Kopiwoda S., Abrams M.J., Darkes M., Robbins R.C., Maecker H.T., Strauss H.W. In vivo detection and imaging of phosphatidylserine expression during programmed cell death. Proc Natl Acad Sci USA 1998; 95(11): 6349–6354, http://dx.doi.org/10.1073/pnas.95.11.6349.
- Kartachova M., van Zandwijk N., Burgers S., van Tinteren H., Verheij M., Valdés Olmos R.A. Prognostic significance of 99mTc Hynic-rh-annexin V scintigraphy during platinum-based chemotherapy in advanced lung cancer. J Clin Oncol 2007; 25(18): 2534–2539, http://dx.doi.org/10.1200/jco.2006.10.1337.
- Kartachova M.S., Valdés Olmos R.A., Haas R.L., Hoebers F.J., Herk M., Verheij M. 99mTc-HYNIC-rh-annexin-V scintigraphy: visual and quantitative evaluation of early treatment-induced apoptosis to predict treatment outcome. Nucl Med Commun 2008; 29(1): 39–44, http://dx.doi.org/10.1097/MNM.0b013e3282f1bc22.
- Hu S., Kiesewetter D.O., Zhu L., Guo N., Gao H., Liu G., Hida N., Lang L., Niu G., Chen X. Longitudinal PET imaging of doxorubicin-induced cell death with 18F-annexin V. Mol Imaging Biol 2012; 14(6): 762–770, http://dx.doi.org/10.1007/s11307-012-0551-5.
- Niu G., Chen X. Apoptosis imaging: beyond annexin V. J Nucl Med 2010; 51(11): 1659–1662, http://dx.doi.org/10.2967/jnumed.110.078584.
- Igarashi K., Kaneda M., Yamaji A., Saido T.C., Kikkawa U., Ono Y., Inoue K., Umeda M. A novel phosphatidylserine-binding peptide motif defined by an anti-idiotypic monoclonal antibody. Localization of phosphatidylserine-specific binding sites on protein kinase C and phosphatidylserine decarboxylase. J Biol Chem 1995; 270(49): 29075–29078, http://dx.doi.org/10.1074/jbc.270.49.29075.
- Xiong C., Brewer K., Song S., Zhang R., Lu W., Wen X., Li C. Peptide-based imaging agents targeting phosphatidylserine for the detection of apoptosis. J Med Chem 2011; 54(6): 1825–1835, http://dx.doi.org/10.1021/jm101477d.
- Song S., Xiong C., Lu W., Ku G., Huang G., Li C. Apoptosis imaging probe predicts early chemotherapy response in preclinical models: a comparative study with 8F-FDG PET. J Nucl Med 2013; 54(1): 104–110, http://dx.doi.org/10.2967/jnumed.112.109397.
- Marconescu A., Thorpe P.E. Coincident exposure of phosphatidylethanolamine and anionic phospholipids on the surface of irradiated cells. Biochim Biophys Acta 2008; 1778(10): 2217–2224, http://dx.doi.org/10.1016/j.bbamem.2008.05.006.
- Zhao M., Li Z., Bugenhagen S. 99mTc-labeled duramycin as a novel phosphatidylethanolamine-binding molecular probe. J Nucl Med 2008; 49(8): 1345–1352, http://dx.doi.org/10.2967/jnumed.107.048603.
- Johnson S.E., Li Z., Liu Y., Moulder J.E., Zhao M. Whole-body imaging of high-dose ionizing irradiation-induced tissue injuries using 99mTc-duramycin. J Nucl Med 2013; 54(8): 1397–1403, http://dx.doi.org/10.2967/jnumed.112.112490.
- Yao S., Hu K., Tang G., Liang X., Du K., Nie D., Jiang S., Zang L. Positron emission tomography imaging of cell death with [18F]FPDuramycin. Apoptosis 2014; 19(5): 841–850, http://dx.doi.org/10.1007/s10495-013-0964-x.
- Luker G.D., Luker K.E. Optical imaging: current applications and future directions. J Nucl Med 2008; 49(1): 1–4, http://dx.doi.org/10.2967/jnumed.107.045799.
- Kumar S., Richards-Kortum R. Optical molecular imaging agents for cancer diagnostics and therapeutics. Nanomedicine 2006; 1: 23–30, http://dx.doi.org/10.2217/17435889.1.1.23.
- Achilefu S. The insatiable quest for near-infrared fluorescent probes for molecular imaging. Angew Chem Int Ed 2010; 49(51): 9816–9818, http://dx.doi.org/10.1002/anie.201005684.
- Petrovsky A., Schellenberger E., Josephson L., Weissleder R., Bogdanov A. Jr. Near-infrared fluorescent imaging of tumor apoptosis. Cancer Res 2003; 63(8): 1936–1942.
- Ntziachristos V., Schellenberger E.A., Ripoll J., Yessayan D., Graves E., Bogdanov A. Jr., et al. Visualization of antitumor treatment by means of fluorescence molecular tomography with an annexin V-Cy5.5 conjugate. Proc Natl Acad Sci USA 2004; 101(33): 12294–12299, http://dx.doi.org/10.1073/pnas.0401137101.
- Hiller K.H., Waller C., Nahrendorf M., Bauer W.R., Jakob P.M. Assessment of cardiovascular apoptosis in the isolated rat heart by magnetic resonance molecular imaging. Mol Imaging 2006; 5(2): 115–121.
- van Tilborg G.A., Mulder W.J., Deckers N., Storm G., Reutelingsperger C.P., Strijkers G.J., Nicolay K. Annexin A5-functionalized bimodal lipid-based contrast agents for the detection of apoptosis. Bioconjug Chem 2006; 17(3): 741–749, http://dx.doi.org/10.1021/bc0600259.
- Zhao M., Beauregard D.A., Loizou L., Davletov B., Brindle K.M. Non-invasive detection of apoptosis using magnetic resonance imaging and a targeted contrast agent. Nat Med 2001; 7(11): 1241–1244, http://dx.doi.org/10.1038/nm1101-1241.
- Krishnan A.S., Neves A.A., de Backer M.M., Hu D.E., Davletov B., Kettunen M.I., Brindle K.M. Detection of cell death in tumors by using MR imaging and a gadolinium-based targeted contrast agent. Radiology 2008; 246(33): 854–862, http://dx.doi.org/10.1148/radiol.2463070471.
- Pop C., Salvesen G.S. Human caspases: activation, specificity, and regulation. J Biol Chem 2009; 284(33): 21777–21781, http://dx.doi.org/10.1074/jbc.R800084200.
- Nicholson D.W., Thornberry N.A. Caspases: killer proteases. Trends Biochem Sci 1997; 22(8): 299–306, http://dx.doi.org/10.1016/S0968-0004(97)01085-2.
- Delgado M.E., Olsson M., Lincoln F.A., Zhivotovsky B., Rehm M. Determining the contributions of caspase-2, caspase-8 and effector caspases to intracellular VDVADase activities during apoptosis initiation and execution. Biochim Biophys Acta 2013; 1833(10): 2279–2292, http://dx.doi.org/10.1016/j.bbamcr.2013.05.025.
- Laussmann M.A., Passante E., Hellwig C.T., Tomiczek B., Flanagan L., Prehn J.H., Huber H.J., Rehm M. Proteasome inhibition can impair caspase-8 activation upon submaximal stimulation of apoptotic tumor necrosis factor-related apoptosis inducing ligand (TRAIL) signaling. J Biol Chem 2012; 287(18): 14402–14411, http://dx.doi.org/10.1074/jbc.M111.304378.
- Yang F., Tu J., Pan J.Q., Luo H.L., Liu Y.H., Wan J., Zhang J., Wei P.F., Jiang T., Chen Y.H., Wang L.P. Light-controlled inhibition of malignant glioma by opsin gene transfer. Cell Death Dis 2013; 4: e893, http://dx.doi.org/10.1038/cddis.2013.425.
- Paulsson J.F., Schultz S.W., Köhler M., Leibiger I., Berggren P.O., Westermark G.T. Real-time monitoring of apoptosis by caspase-3-like protease induced FRET reduction triggered by amyloid aggregation. Exp Diabetes Res 2008; 2008: 865850, http://dx.doi.org/10.1155/2008/865850.
- Li I.T., Pham E., Truong K. Protein biosensors based on the principle of fluorescence resonance energy transfer for monitoring cellular dynamics. Biotechnol Lett 2006; 28(24): 1971–1982, http://dx.doi.org/10.1007/s10529-006-9193-5.
- Keese M., Offterdinger M., Tischer C., Girod A., Lommerse P.H., Yagublu V., Magdeburg R., Bastiaens P.I. Quantitative imaging of apoptosis commitment in colorectal tumor cells. Differentiation 2007; 75(9): 809–818, http://dx.doi.org/10.1111/j.1432-0436.2007.00186.x.
- Keese M., Yagublu V., Schwenke K., Post S., Bastiaens P. Fluorescence lifetime imaging microscopy of chemotherapy-induced apoptosis resistance in a syngenic mouse tumor model. Int J Cancer 2010; 126(1): 104–113, http://dx.doi.org/10.1002/ijc.24730.
- Kawai H., Suzuki T., Kobayashi T., Sakurai H., Ohata H., Honda K., Momose K., Namekata I., Tanaka H., Shigenobu K., Nakamura R., Hayakawa T., Kawanishi T. Simultaneous real-time detection of initiator- and effector-caspase activation by double fluorescence resonance energy transfer analysis. J Pharmacol Sci 2005; 93(7): 361–368, http://dx.doi.org/10.1254/jphs.fp0040592.
- Seervi M., Sobhan P.K., Mathew K.A., Joseph J., Pillai P.R., Santhoshkumar T.R. A high-throughput image-based screen for the identification of Bax/Bak-independent caspase activators against drug-resistant cancer cells. Apoptosis 2014; 19(1): 269–284, http://dx.doi.org/10.1007/s10495-013-0921-8.
- Rehm M., Dussmann H., Janicke R.U., Tavare J.M., Kogel D., Prehn J.H. Single-cell fluorescence resonance energy transfer analysis demonstrates that caspase activation during apoptosis is a rapid process. Role of caspase-3. J Biol Chem 2002; 277(27): 24506–24514, http://dx.doi.org/10.1074/jbc.m110789200.
- Wu Y., Xing D., Luo S., Tang Y., Chen Q. Detection of caspase-3 activation in single cells by fluorescence resonance energy transfer during photodynamic therapy induced apoptosis. Cancer Lett 2006; 235(2): 239–247, http://dx.doi.org/10.1016/j.canlet.2005.04.036.
- Janssen A., Beerling E., Medema R., van Rheenen J. Intravital FRET imaging of tumor cell viability and mitosis during chemotherapy. PLoS One 2013; 8(5): e64029, http://dx.doi.org/10.1371/journal.pone.0064029.
- Joseph J., Seervi M., Sobhan P.K., Retnabai S.T. High throughput ratio imaging to profile caspase activity: potential application in multiparameter high content apoptosis analysis and drug screening. PLoS One 2011; 6(5): e20114, http://dx.doi.org/10.1371/journal.pone.0020114.
- Pan W., Qu J., Chen T., Sun L., Qi J. FLIM and emission spectral analysis of caspase-3 activation inside single living cell during anticancer drug-induced cell death. Eur Biophys J 2009; 38(4): 447–456, http://dx.doi.org/10.1007/s00249-008-0390-0.
- Wang L., Chen T., Qu J., Wei X. Quantitative analysis of caspase-3 activation by fitting fluorescence emission spectra in living cells. Micron 2009; 40(8): 811–820, http://dx.doi.org/10.1016/j.micron.2009.07.001.
- Zhou F., Xing D., Wu S., Chen W.R. Intravital imaging of tumor apoptosis with FRET probes during tumor therapy. Mol Imaging Biol 2010; 12(1): 63–70, http://dx.doi.org/10.1007/s11307-009-0235-y.
- Wang F., Chen T.S., Xing D., Wang J.J., Wu Y.X. Measuring dynamics of caspase-3 activity in living cells using FRET technique during apoptosis induced by high fluence low-power laser irradiation. Lasers Surg Med 2005; 36(1): 2–7, http://dx.doi.org/10.1002/lsm.20130.
- Kominami K., Nagai T., Sawasaki T., Tsujimura Y., Yashima K., Sunaga Y., Tsuchimochi M., Nishimura J., Chiba K., Nakabayashi J., Koyamada K., Endo Y., Yokota H., Miyawaki A., Manabe N., Sakamaki K. In vivo imaging of hierarchical spatiotemporal activation of caspase-8 during apoptosis. PLoS One 2012; 7(11): e50218, http://dx.doi.org/10.1371/journal.pone.0050218.
- Pepperkok R., Squire A., Geley S., Bastiaens P.I. Simultaneous detection of multiple green fluorescent proteins in live cells by fluorescence lifetime imaging microscopy. Curr Biol 1999; 9(5): 269–272, http://dx.doi.org/10.1016/s0960-9822(99)80117-1.
- van Kuppeveld F.J., Melchers W.J., Willems P.H., Gadella T.W. Homomultimerization of the coxsackievirus 2B protein in living cells visualized by fluorescence resonance energy transfer microscopy. J Virol 2002; 76(18): 9446–9456, http://dx.doi.org/10.1128/jvi.76.18.9446-9456.2002.
- Chen N.T., Wu C.Y., Chung C.Y., Hwu Y., Cheng S.H., Mou C.Y., Lo L.W. Probing the dynamics of doxorubicin-DNA intercalation during the initial activation of apoptosis by fluorescence lifetime imaging microscopy (FLIM). PLoS One 2012; 7(9): e44947, http://dx.doi.org/10.1371/journal.pone.0044947.
- Verveer P.J., Squire A., Bastiaens P.I. Global analysis of fluorescence lifetime imaging microscopy data. Biophys J 2000; 78(4): 2127–2137, http://dx.doi.org/10.1016/s0006-3495(00)76759-2.
- Angres B., Steuer H., Weber P., Wagner M., Schneckenburger H. A membrane-bound FRET-based caspase sensor for detection of apoptosis using fluorescence lifetime and total internal reflection microscopy. Cytometry A 2009; 75(5): 420–427, http://dx.doi.org/10.1002/cyto.a.20698.
- Weber P., Schickinger S., Wagner M., Angres B., Bruns T., Schneckenburger H. Monitoring of apoptosis in 3D cell cultures by FRET and light sheet fluorescence microscopy. Int J Mol Sci 2015; 16(3): 5375–5385, http://dx.doi.org/10.3390/ijms16035375.
- Bozza W.P., Di X., Takeda K., Rivera Rosado L.A., Pariser S., Zhang B. The use of a stably expressed FRET biosensor for determining the potency of cancer drugs. PLoS One 2014; 9(9): e107010, http://dx.doi.org/10.1371/journal.pone.0107010.
- Savitsky A.P., Rusanov A.L., Zherdeva V.V., Gorodnicheva T.V., Khrenova M.G., Nemukhin A.V. FLIM-FRET imaging of caspase-3 activity in live cells using pair of red fluorescent proteins. Theranostics 2012; 2(2): 215–226, http://dx.doi.org/10.7150/thno.3885.
- Rusanov A.L., Ivashina T.V., Vinokurov L.M., Fiks I.I., Orlova A.G., Turchin I.V., Meerovich I.G., Zherdeva V.V., Savitsky A.P. Lifetime imaging of FRET between red fluorescent proteins. J Biophotonics 2010; 3(12): 774–783, http://dx.doi.org/10.1002/jbio.201000065.
- Shcherbo D., Souslova E.A., Goedhart J., Chepurnykh T.V., Gaintzeva A., Shemiakina I.I., Gadella T.W., Lukyanov S., Chudakov D.M. Practical and reliable FRET/FLIM pair of fluorescent proteins. BMC Biotechnol 2009; 9: 24, http://dx.doi.org/10.1186/1472-6750-9-24.
- Shcherbo D., Shemiakina I.I., Ryabova A.V., Luker K.E., Schmidt B.T., Souslova E.A., Gorodnicheva T.V., Strukova L., Shidlovskiy K.M., Britanova O.V., Zaraysky A.G., Lukyanov K.A., Loschenov V.B., Luker G.D., Chudakov D.M. Near-infrared fluorescent proteins. Nat Methods 2010; 7(10): 827–829, http://dx.doi.org/10.1038/nmeth.1501.
- Shu X., Royant A., Lin M.Z., Aguilera T.A., Lev-Ram V., Steinbach P.A., Tsien R.Y. Mammalian expression of infrared fluorescent proteins engineered from a bacterial phytochrome. Science 2009; 324(5928): 804–807, http://dx.doi.org/10.1126/science.1168683.
- Shcherbakova D.M., Verkhusha V.V. Near-infrared fluorescent proteins for multicolor in vivo imaging. Nat Methods 2013; 10(8): 751–754, http://dx.doi.org/10.1038/nmeth.2521.
- Subach O.M., Gundorov I.S., Yoshimura M., Subach F.V., Zhang J., Grüenwald D., Souslova E.A., Chudakov D.M., Verkhusha V.V. Conversion of red fluorescent protein into a bright blue probe. Chem Biol 2008; 15(10): 1116–1124, http://dx.doi.org/10.1016/j.chembiol.2008.08.006.
- Filonov G.S., Piatkevich K.D., Ting L.M., Zhang J., Kim K., Verkhusha V.V. Bright and stable near-infrared fluorescent protein for in vivo imaging. Nat Biotechnol 2011; 29(8): 757–761, http://dx.doi.org/10.1038/nbt.1918.
- Rajoria S., Zhao L., Intes X., Barroso M. FLIM-FRET for cancer applications. Curr Mol Imaging 2014; 3(2): 144–161, http://dx.doi.org/10.2174/2211555203666141117221111.
- Nguyen Q.D., Lavdas I., Gubbins J., Smith G., Fortt R., Carroll L.S., Graham M.A., Aboagye E.O. Temporal and spatial evolution of therapy-induced tumor apoptosis detected by caspase-3-selective molecular imaging. Clin Cancer Res 2013; 19: 3914–3924, http://dx.doi.org/10.1158/1078-0432.CCR-12-3814.
- Challapalli A., Kenny L.M., Hallett W.A., Kozlowski K., Tomasi G., Gudi M., Al-Nahhas A., Coombes R.C., Aboagye E.O. 18F-ICMT-11, a caspase-3-specific PET tracer for apoptosis: biodistribution and radiation dosimetry. J Nucl Med 2013; 54(9): 1551–1556, http://dx.doi.org/10.2967/jnumed.112.118760.
- Susin S.A., Lorenzo H.K., Zamzami N., Marzo I., Snow B.E., Brothers G.M., Mangion J., Jacotot E., Costantini P., Loeffler M., Larochette N., Goodlett D.R., Aebersold R., Siderovski D.P., Penninger J.M., Kroemer G. Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 1999; 397(6718): 441–446, http://dx.doi.org/10.1038/17135.
- Maltese W.A., Overmeyer J.H. Non-apoptotic cell death associated with perturbations of macropinocytosis. Front Physiol 2015; 6: 38, http://dx.doi.org/10.3389/fphys.2015.00038.
- Susin S.A., Zamzami N., Kroemer G. Mitochondria as regulators of apoptosis: doubt no more. Biochim Biophys Acta 1998; 1366(1–2): 151–165, http://dx.doi.org/10.1016/S0005-2728(98)00110-8.
- Green D.R., Kroemer G. The pathophysiology of mitochondrial cell death. Science 2004; 305(5684): 626–629, http://dx.doi.org/10.1126/science.1099320.
- Bouchier-Hayes L., Muсoz-Pinedo C., Connell S., Green D.R. Measuring apoptosis at the single cell level. Methods 2008; 44(3): 222–228, http://dx.doi.org/10.1016/j.ymeth.2007.11.007.
- Madar I., Ravert H., Nelkin B., Abro M., Pomper M., Dannals R., Frost J.J. Characterization of membrane potential-dependent uptake of the novel PET tracer 18F-fluorobenzyl triphenylphosphonium cation. Eur J Nucl Med Mol Imaging 2007; 34(12): 2057–2065, http://dx.doi.org/10.1007/s00259-007-0500-8.
- Madar I., Huang Y., Ravert H., Dalrymple S.L., Davidson N.E., Isaacs J.T., Dannals R.F., Frost J.J. Detection and quantification of the evolution dynamics of apoptosis using the PET voltage sensor F-18-fluorobenzyl triphenyl phosphonium. J Nucl Med 2009; 50: 774–780, http://dx.doi.org/10.2967/jnumed.108.061283.
- Bortner C.D., Oldenburg N.B., Cidlowski J.A. The role of DNA fragmentation in apoptosis. Trends Cell Biol 1995; 5(1): 21–26, http://dx.doi.org/10.1016/S0962-8924(00)88932-1.
- Gorczyca W., Gong J., Darzynkiewicz Z. Detection of DNA strand breaks in individual apoptotic cells by the in situ terminal deoxynucleotidyl transferase and nick translation assays. Cancer Res 1993; 53: 1945–1951.
- Negoescu A., Guillermet C., Lorimier P., Brambilla E., Labat-Moleur F. Importance of DNA fragmentation in apoptosis with regard to TUNEL specificity. Biomed Pharmacother 1998; 52(6): 252–258, http://dx.doi.org/10.1016/S0753-3322(98)80010-3.
- Diaconu C.C., Szathmári M., Kéri G., Venetianer A. Apoptosis is induced in both drug-sensitive and multidrug-resistant hepatoma cells by somatostatin analogue TT-232. Br J Cancer 1999; 80(8): 1197–1203, http://dx.doi.org/10.1038/sj.bjc.6690486.
- Naruse I., Keino H., Kawarada Y. Antibody against single-stranded DNA detects both programmed cell death and drug-induced apoptosis. Histochemistry 1994; 101(1): 73–78, http://dx.doi.org/10.1007/bf00315834.
- Cornelissen B., Kersemans V., Darbar S., Thompson J., Shah K., Sleeth K., Hill M.A., Vallis K.A. Imaging DNA damage in vivo using gammaH2AX-targeted immunoconjugates. Cancer Res 2011; 71(13): 4539–4549, http://dx.doi.org/10.1158/0008-5472.CAN-10-4587.
- Porębska I., Kosacka M., Sobańska E., Wyrodek E., Jankowska R. Comparative expression of apoptotic markers in lung adenocarcinoma and squamous cell carcinoma. Adv Exp Med Biol 2015, http://dx.doi.org/10.1007/5584_2015_121. [Epub ahead of print].
- Wu Y., Kim S.G., Xing D., Liu L., Park J.C., Chen T., Chen W.R. Fluorescence resonance energy transfer analysis of bid activation in living cells during ultraviolet-induced apoptosis. Acta Biochim Biophys Sin (Shanghai) 2007; 39(1): 37–45, http://dx.doi.org/10.1111/j.1745-7270.2007.00246.x.
- Louie A. Multimodality imaging probes: design and challenges. Chem Rev 2010; 110(5): 3146–3195, http://dx.doi.org/10.1021/cr9003538.
- Sapsford K.E., Algar W.R., Berti L., Gemmill K.B., Casey B.J., Oh E., Stewart M.H., Medintz I.L. Functionalizing nanoparticles with biological molecules: developing chemistries that facilitate nanotechnology. Chem Rev 2013; 113(3): 1904–2074, http://dx.doi.org/10.1021/cr300143v.
- Xing Y., Zhao J., Conti P.S., Chen K. Radiolabeled nanoparticles for multimodality tumor imaging. Theranostics 2014; 4: 290–306, http://dx.doi.org/10.7150/thno.7341.