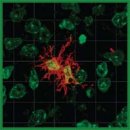
Immunohistochemical Detection of Microglia Using Iba-1 as a Marker in Health and Disease
The aim of the study was to evaluate the possibilities and limitations of using the immunohistochemical detection of Iba-1 protein for morphofunctional analysis of microglia under different conditions.
Materials and Methods. In the study we used brain samples from Wistar rats at different ages: 7 days of postnatal development (n=18), 14 days of postnatal development (n=18), 4–6 months (n=22); the brain samples from 3–6-month-old SHR (spontaneously hypertensive rats) male rats (n=4); human cerebral cortex samples (n=10). Rabbit polyclonal antibodies against calcium-binding protein Iba-1 (Biocare Medical, USA) were used to detect microglia immunohistochemically followed by light and confocal laser microscopy.
A previously developed original technique was applied to simultaneously detect microglial cells and amyloid plaques.
Results. Iba-1 protein in microglial cells was shown to be present in grey and white matter in all cerebral regions under study. Microglial cells in different regions were noted to be characterized by pronounced structural and functional features when stained for Iba-1. In addition to microglial cells, in the rat brain there were found other Iba-1-immunopositive macrophages, which were tissue cerebral macrophages differing from microgliocytes by specific morphology and localization. Iba-1 protein was demonstrated to be present in microglial cells in all investigated stages of postnatal development that makes Iba-1 an appropriate marker for comparative ontogenetical studies of microglial cells. The localization of Iba-1 in the bodies and processes of microgliocytes enables to more clearly identify the complex ramified cell morphology and make three-dimensional reconstructions. In the brain of spontaneously hypertensive rats (SHR line rats) the Iba-1-immunopositive microglia were shown to have a number of structural and functional features indicating the moderate activation of microglia. 48 h after ischemic injury, an anomalously great number of large Iba-1-immunopositive cells with amoeboid morphology were detected in the gray matter of the striatum in an ipsilateral hemisphere near the injured area in the rat brain. Microglial cells in human cerebral cortex were noted to be localized in the majority of the detected amyloid plaques and characterized by amoeboid morphology indicating their severe activation.
Conclusion. The presented findings suggest Iba-1 protein to be a reliable and universal microglial marker. The immunohistochemical detection of the protein enables to identify and analyze qualitatively and quantitatively microglial cells in different brain regions in human and laboratoryanimals under normal and pathological conditions. The limitations of using Iba-1 as a marker of microglia include the inability to determine the microglial activation vector and the difficulty in distinguishing between microglia and infiltrating macrophages of the brain in pathology. In such cases, it is necessary to improve the technology for detecting microglia that can be based on using a multi-marker analysis.
- Adamu A., Li S., Gao F., Xue G. The role of neuroinflammation in neurodegenerative diseases: current understanding and future therapeutic targets. Front Aging Neurosci 2024; 16: 1347987, https://doi.org/10.3389/fnagi.2024.1347987.
- Muzio L., Viotti A., Martino G. Microglia in neuroinflammation and neurodegeneration: from understanding to therapy. Front Neurosci 2021; 15: 742065, https://doi.org/10.3389/fnins.2021.742065.
- Healy L.M., Zia S., Plemel J.R. Towards a definition of microglia heterogeneity. Commun Biol 2022; 5(1): 1114, https://doi.org/10.1038/s42003-022-04081-6.
- Malinovskaya N.A., Frolova O.V., Shishelova K.O., Panina Yu.A. Current methods for the isolation and cultivation of microglia (review). Sovremennye tehnologii v medicine 2021; 13(6): 89, https://doi.org/10.17691/stm2021.13.6.10.
- Jurga A.M., Paleczna M., Kuter K.Z. Overview of general and discriminating markers of differential microglia phenotypes. Front Cell Neurosci 2020; 14: 198, https://doi.org/10.3389/fncel.2020.00198.
- Cserép C., Pósfai B., Dénes Á. Shaping neuronal fate: functional heterogeneity of direct microglia-neuron interactions. Neuron 2021; 109(2): 222–240, https://doi.org/10.1016/j.neuron.2020.11.007.
- Masuda T., Sankowski R., Staszewski O., Prinz M. Microglia heterogeneity in the single-cell era. Cell Rep 2020; 30(5): 1271–1281, https://doi.org/10.1016/j.celrep.2020.01.010.
- Galatro T.F., Holtman I.R., Lerario A.M., Vainchtein I.D., Brouwer N., Sola P.R., Veras M.M., Pereira T.F., Leite R.E.P., Möller T., Wes P.D., Sogayar M.C., Laman J.D., den Dunnen W., Pasqualucci C.A., Oba-Shinjo S.M., Boddeke E.W.G.M., Marie S.K.N., Eggen B.J.L. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat Neurosci 2017; 20(8): 1162–1171, https://doi.org/10.1038/nn.4597.
- Tan Y.L., Yuan Y., Tian L. Microglial regional heterogeneity and its role in the brain. Mol Psychiatry 2020; 25(2): 351–367, https://doi.org/10.1038/s41380-019-0609-8.
- Hickman S.E., Kingery N.D., Ohsumi T.K., Borowsky M.L., Wang L.C., Means T.K., El Khoury J. The microglial sensome revealed by direct RNA sequencing. Nat Neurosci 2013; 16(12): 1896–1905, https://doi.org/10.1038/nn.3554.
- Mildner A., Huang H., Radke J., Stenzel W., Priller J. P2Y12 receptor is expressed on human microglia under physiological conditions throughout development and is sensitive to neuroinflammatory diseases. Glia 2017; 65(2): 375–387, https://doi.org/10.1002/glia.23097.
- Zia S., Rawji K.S., Michaels N.J., Burr M., Kerr B.J., Healy L.M., Plemel J.R. Microglia diversity in health and multiple sclerosis. Front Immunol 2020; 11: 588021, https://doi.org/10.3389/fimmu.2020.588021.
- Fumagalli L., Nazlie Mohebiany A., Premereur J., Polanco Miquel P., Bijnens B., Van de Walle P., Fattorelli N., Mancuso R. Microglia heterogeneity, modeling and cell-state annotation in development and neurodegeneration. Nat Neurosci 2025; 28(7): 1381–1392, https://doi.org/10.1038/s41593-025-01931-4.
- Kanazawa H., Ohsawa K., Sasaki Y., Kohsaka S., Imai Y. Macrophage/microglia-specific protein Iba1 enhances membrane ruffling and Rac activation via phospholipase C-gamma-dependent pathway. J Biol Chem 2002; 277(22): 20026–20032, https://doi.org/10.1074/jbc.M109218200.
- Kolos E.A., Korzhevskii D.E. Spinal cord microglia in health and disease. Acta Naturae 2020; 12(1): 4–17, https://doi.org/10.32607/actanaturae.10934.
- Dyban A.P., Puchkov B.F., Baranov V.S., Samoshkina N.A., Chebotar N.A. Laboratornye mlekopitayushchie: mysh' Mus musculus, krysa Rattus norvegicus, krolik Oryctolagus cuniculus, khomyachok Cricetus griseous. V kn.: Ob"ekty biologii razvitiya [Laboratory mammals: mouse Mus musculus, rat Rattus norvegicus, rabbit Oryctolagus cuniculus, hamster Cricetus griseous. In: Objects of developmental biology]. Moscow: Nauka; 1975; p. 505–566.
- Korzhevskii D.E., Kirik O.V., Vlasov T.D. Appearance of stellate smooth muscle cells in the rat brain after transitory focal ischemia. Morphology 2013; 143(1): 073–075.
- Guselnikova V.V., Antipova M.V., Fedorova E.A., Safray A.E., Rukavishnikova A.A., Mikhailova E.V., Korzhevskii D.E. Distinctive features of histochemical and immunohistochemical techniques for amyloid plaque detection in the human cerebral cortex. Journal of Anatomy and Histopathology 2019; 8(2): 91–99, https://doi.org/10.18499/2225-7357-2019-8-2-91-99.
- Korzhevskii D.E., Sukhorukova Y.G., Gilerovich Y.G., Petrova Y.S., Kirik O.V., Grigoriev I.P. Advantages and disadvantages of zinc-ethanol-formaldehyde as a fixative for immunocytochemical studies and confocal laser microscopy. Morphology 2013; 143(2): 081–085.
- Nosova O.I., Guselnikovа V.V., Korzhevskii D.E. Light microscopy approach for simultaneously identification of glial cells and amyloid plaques. Tsitologiya 2021; 63(6): 557–567, https://doi.org/10.31857/S0041377121060092.
- Korzhevskii D.E., Kirik O., Sukhorukova E. Immunocytochemistry of microglial cells. In: Merigh A., Lossi L. (editors). Immunocytochemistry and related techniques. Neuromethods. Vol. 101. Humana Press, New York; 2015, https://doi.org/10.1007/978-1-4939-2313-7_12.
- Bennett M.L., Bennett F.C., Liddelow S.A., Ajami B., Zamanian J.L., Fernhoff N.B., Mulinyawe S.B., Bohlen C.J., Adil A., Tucker A., Weissman I.L., Chang E.F., Li G., Grant G.A., Hayden Gephart M.G., Barres B.A. New tools for studying microglia in the mouse and human CNS. Proc Natl Acad Sci U S A 2016; 113(12): E1738–E1746, https://doi.org/10.1073/pnas.1525528113.
- Lier J., Streit W.J., Bechmann I. Beyond activation: characterizing microglial functional phenotypes. Cells 2021; 10(9): 2236, https://doi.org/10.3390/cells10092236.
- Hirasawa T., Ohsawa K., Imai Y., Ondo Y., Akazawa C., Uchino S., Kohsaka S. Visualization of microglia in living tissues using Iba1-EGFP transgenic mice. J Neurosci Res 2005; 81(3): 357–362, https://doi.org/10.1002/jnr.20480.
- Shui X., Chen J., Fu Z., Zhu H., Tao H., Li Z. Microglia in ischemic stroke: pathogenesis insights and therapeutic challenges. J Inflamm Res 2024; 17: 3335–3352, https://doi.org/10.2147/JIR.S461795.
- Taylor R.A., Sansing L.H. Microglial responses after ischemic stroke and intracerebral hemorrhage. Clin Dev Immunol 2013; 2013: 746068, https://doi.org/10.1155/2013/746068.
- Korzhevskii D.E., Kirik O.V., Sukhorukova Ye.G., Vlasov T.D. Structural organization of striatal microgliocytes following focal ischemia. Morphology 2012; 141(2): 28–32.
- Murata Y., Sugimoto K., Yang C., Harada K., Gono R., Harada T., Miyashita Y., Higashisaka K., Katada R., Tanaka J., Matsumoto H. Activated microglia-derived macrophage-like cells exacerbate brain edema after ischemic stroke correlate with astrocytic expression of aquaporin-4 and interleukin-1 alpha release. Neurochem Int 2020; 140: 104848, https://doi.org/10.1016/j.neuint.2020.104848.
- Hansen D.V., Hanson J.E., Sheng M. Microglia in Alzheimer's disease. J Cell Biol 2018; 217(2): 459–472, https://doi.org/10.1083/jcb.201709069.
- Grubman A., Choo X.Y., Chew G., Ouyang J.F., Sun G., Croft N.P., Rossello F.J., Simmons R., Buckberry S., Landin D.V., Pflueger J., Vandekolk T.H., Abay Z., Zhou Y., Liu X., Chen J., Larcombe M., Haynes J.M., McLean C., Williams S., Chai S.Y., Wilson T., Lister R., Pouton C.W., Purcell A.W., Rackham O.J.L., Petretto E., Polo J.M. Transcriptional signature in microglia associated with Aβ plaque phagocytosis. Nat Commun 2021; 12(1): 3015, https://doi.org/10.1038/s41467-021-23111-1.
- Miao J., Ma H., Yang Y., Liao Y., Lin C., Zheng J., Yu M., Lan J. Microglia in Alzheimer's disease: pathogenesis, mechanisms, and therapeutic potentials. Front Aging Neurosci 2023; 15: 1201982, https://doi.org/10.3389/fnagi.2023.1201982.
- Bubnova S., Guryanov A., Konkina D., Karapetian V., Kaldybaev l., Pavlova D., Shorina V., Sigalov M., Сhen C. Evaluation of the levels of inflammatory proteins, such as Iba1 and GFAP, in brain tissues of patients with Alzheimer’s disease by western blot analysis. Arzneimitteltherapie 2025; 1385–1397. URL: https://arzneimitteltherapie.com/archives/1520.
- Lier J., Winter K., Bleher J., Grammig J., Mueller W.C., Streit W., Bechmann I. Loss of Iba1-expression in brains from individuals with obesity and hepatic dysfunction. Brain Res 2019; 1710: 220–229, https://doi.org/10.1016/j.brainres.2019.01.006.
- Imai Y., Ibata I., Ito D., Ohsawa K., Kohsaka S. A novel gene Iba1 in the major histocompatibility complex class III region encoding an EF hand protein expressed in a monocytic lineage. Biochem Biophys Res Commun 1996; 224(3): 855–862, https://doi.org/10.1006/bbrc.1996.1112.
- Hollopeter G., Jantzen H.M., Vincent D., Li G., England L., Ramakrishnan V., Yang R.B., Nurden P., Nurden A., Julius D., Conley P.B. Identification of the platelet ADP receptor targeted by antithrombotic drugs. Nature 2001; 409(6817): 202–207, https://doi.org/10.1038/35051599.
- Butovsky O., Jedrychowski M.P., Moore C.S., Cialic R., Lanser A.J., Gabriely G., Koeglsperger T., Dake B., Wu P.M., Doykan C.E., Fanek Z., Liu L., Chen Z., Rothstein J.D., Ransohoff R.M., Gygi S.P., Antel J.P., Weiner H.L. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat Neurosci 2014; 17(1): 131–143, https://doi.org/10.1038/nn.3599.
- Maeda J., Minamihisamatsu T., Shimojo M., Zhou X., Ono M., Matsuba Y., Ji B., Ishii H., Ogawa M., Akatsu H., Kaneda D., Hashizume Y., Robinson J.L., Lee V.M., Saito T., Saido T.C., Trojanowski J.Q., Zhang M.R., Suhara T., Higuchi M., Sahara N. Distinct microglial response against Alzheimer's amyloid and tau pathologies characterized by P2Y12 receptor. Brain Commun 2021; 3(1): fcab011, https://doi.org/10.1093/braincomms/fcab011.
- Amadio S., Parisi C., Montilli C., Carrubba A.S., Apolloni S., Volonté C. P2Y(12) receptor on the verge of a neuroinflammatory breakdown. Mediators Inflamm 2014; 2014: 975849, https://doi.org/10.1155/2014/975849.










