16S rRNA Metabarcoding Used to Identify the Etiology of Infective Endocarditis Agents
The aim of the study was to estimate the efficiency of identifying bacterial agents in native cardiac valves affected by infective endocarditis using 16S rRNA metabarcoding.
Materials and Methods. The study material involved 20 native cardiac valve samples from 16 patients. Sequencing was carried out in the Center for Collective Use “Genomics” (Institute of Chemical Biology and Fundamental Medicine, Siberian Branch of the Russian Academy of Sciences, Russia) on a sequenator MiSeq (Illumina, USA) using MiSeq Reagent Kit v3 (2×300 bp; Illumina, USA).
Results. The study revealed major microorganisms (their portion in the sample under study was over 5% from all identified bacterial agents) belonging to Streptococcus (40% of all valves studied), Sphingomonas (35%), Pseudomonas (35%), Roseateles (25%), Phyllobacterium (25%), and Enterococcus (15%). Moreover, the studied valves were found to have rare cases of Ralstonia pickettii, as well as Bacillus and Klebsiella representatives.
Conclusion. The findings demonstrated the efficiency of 16S rRNA metabarcoding in identifying bacterial agents compared to routine noninvasive diagnostic methods. Native cardiac valves isolated from the patients with infective endocarditis were characterized by numerous opportunistic pathogenic bacteria in negative blood cultures.
Introduction
Infective endocarditis (IE) is an inflammatory condition of primarily bacterial origin characterized by agents localized on native cardiac valval structures, endocardium, as well as on prosthetic cardiac valves and pacemakers [1]. Epidemiological studies have demonstrated an increase in IE incidence rate and the growth of fatal cases due to the complications related to the pathological condition [2, 3]. Agent identification is a crucially important aspect for IE diagnosis; it has an impact on antibiotic therapy and recurrent bacteremia risk in patients with cardiac valve prostheses [4]. The gold standard of identifying IE etiology is considered to be a microbiological blood culture or cardiac valval tissue excised. However, it should be noted that in most cases the cultures appear to be negative. In this situation, according to applicable clinical guidelines [5], it is possible to use molecular genetic methods (e.g., polymerase chain reaction) [6]. Preoperative antibiotic therapy administered in an inpatient department implies difficulties in detecting the agents using standard methods; therefore, there is an increasing demand for more accurate diagnosis and identification of bacterial agents in order to decrease IE mortality [7].
Currently, one of the promising methods is next-generation sequencing (NGS), which enables to identify pathogens in cases when standard molecular genetic tests yield no positive result [8]. It should be mentioned that the sensitivity and specificity of the method are higher for explanted tissues compared to peripheral blood [9]. At the initial stage of metagenomic 16S rRNA sequencing, there is the amplification of variable parts using specific primers followed by sequencing the resulted product that enables to identify a bacteriological agent, its genus, species. 16S rRNA gene, which encodes the 16S ribosome subunit, has been found in all bacteria, it includes highly conserved regions with variable sequences [10]. The unquestionable advantage of the method is detecting nonviable microorganisms by a bacterial DNA, as well as the bacterial agents in patients after antibiotic therapy [11].
The aim of the study was to estimate the efficiency of identifying bacterial agents in native cardiac valves affected by infective endocarditis using 16S rRNA metabarcoding.
Material and Methods
Study group. Study material involved 20 native cardiac valve specimens affected by IE, which were operatively taken from 16 cardiosurgical patients. It should be noted that the most cases were left-sided IE, while right-sided IE was found in 1 patient.
Clinical patient profiles:
male gender — 12 patients (75%);
mean age (Me [Q1; Q3]) — 57.0 [35.0; 63.0] years;
body mass index (mean±SD) — 24.63±3.95;
left ventricular ejection fraction (mean±SD) — (63.64±9.50)%;
pulmonary hypertension — 10 patients (62.5%);
hypertensive disease — 7 patients (43.75%);
chronic cardiac failure — 16 patients (100%);
atrial fibrillation — 3 patients (18.75%);
acute cerebrovascular disease — 4 patients (25%);
chronic renal failure — 8 patients (50%);
chronic obstructive pulmonary disease — 4 patients (25%);
antibiotics taken prior to admission — 12 patients (75%);
involved valve: aortal, mitral and tricuspid — 9 (45%) patients, 10 (50%) patients, and 1 (5%) patient, respectively.
Laboratory findings (Me [Q1; Q3]):
leukocytes — 7.20 [5.40; 9.30]·109/L;
neutrophils — 4.40 [2.90; 5.85]·109/L;
C-reactive protein — 4.80 [2.00; 16.40] mg/L;
erythrocyte sedimentation rate — 18.0 [5.50; 51.50] mm/h.
The study was carried out in accordance with the standards of Good Clinical Practice and Declaration of Helsinki (2024). All participants signed the informed consent to be involved in the research. The present study was approved by a local Ethics Committee of Research Institute for Complex Issues of Cardiovascular Diseases (protocol No.1 dated January 26, 2024).
Infective endocarditis diagnosis was made based on clinical, microbiological, and laboratory data, and echocardiography (echoCG) findings. According to echoCG, all patients involved in the study, had movable vegetations of the valves. Additionally, the diagnosis was verified in accordance with Duke international criteria. A week prior to the surgery, the patients under study had a peripheral blood test for microbiological testing in three repeats, the interval being 6 h. The histological Gram staining was performed using a commercial kit (Abcam, Great Britain).
16S rRNA metabarcoding. DNA was isolated from the explanted valval specimens using HostZERO Microbial DNA Kit (Zymo Research, USA). V3–V4 region of 16S rRNA gene was amplified using the primers: 343F (5’-CTCCTACGGRRSGCAGCAG-3’) and 806R (5’-GGACTACNVGGGTWTCTAAT-3’) containing the adapter sequences (Illumina, USA), a linker and a barcode [12]. The amplification was performed in the reaction mixture, 20 µl (in three repetitions) containing Hot Start Taq DNA polymerase, 0.4 U (Biolabmix, Russia) and single HS Taq PCR-buffer (Biolabmix, Russia); 0.2 µmol of forward and reverse primers; DNA, 10 ng; MgCl2, 2.3 mM (Biolabmix, Russia), and dNTP, 0.2 mM (Life Technologies, USA). The number of cycles was selected for each DNA sample, performing PCR with fluorescent signal detection in a real-time mode on a DNA-amplifier CFX-96 (Bio-Rad, USA). As an intercalating fluorophore we used a dye — Eva488 (Lumiprobe, USA). A cycle selection criterion was a fluorescent signal located at a linear logarithmic growth phase. In case of low PCR efficiency, the maximal number of cycles was 34. The target product yield was analyzed on MultiNA using 12000 DNA Base Pair Kit (Shimadzu, Japan). PCR was performed as follows: 5 min at 95°С; then up to 40 cycles — 15 s at 95°С, 15 s at 62°С, 30 s at 72°С; the last stage — 5 min at 72°С. Amplicons (by 200 ng each) were united and cleaned using magnetic particles AMPure XP Beads (Beckman Coulter, USA) according to a manufacturer’s instruction.
Sequencing was carried out in the Center for Collective Use “Genomics” (Institute of Chemical Biology and Fundamental Medicine, Siberian Branch of the Russian Academy of Sciences) on a sequenator MiSeq (Illumina, USA) using MiSeq Reagent Kit v3 (2×300 bp; Illumina, USA). The number of reads obtained for each sample was determined using the program Seqkit [13]. The quality was studied using FastQC [14]. Pair sequences were analyzed using UPARSE-scripts [15] and Usearch v. 11.0.667 [16]. Bioinformation processing included the overlapping of paired reads, the filtration by quality and length, the consideration of similar sequences, dropping of singletons, deletion of chimeras, and obtaining OTU using a clustering algorithm UPARSE [17]. The taxonomic affiliation of OTU sequences was determined using SINTAX [18] and the reference base 16S RDP training set v. 19 [19].
Statistical data processing. The data werestatistically processed using the program Prism 8 (GraphPad Software, USA). Quantitative results were represented as arithmetic mean (М) and standard deviation (SD), as well as median (Me), lower and higher quartiles [Q1; Q3].
Results
A microbiological blood test carried out in accordance with a standard protocol and described in clinical guidelines [5] enabled to state IE etiology in only 25% cases, when IE agents were gram-positive bacteria. Histological Gram staining of the incised cusps of native cardiac valves succeeded in identifying the bacteriological agents in 45% cases (see the Table). In most cases, 16S rRNA metabarcoding revealed major microorganisms (their portion in the samples under study was over 5% of all identified bacterial agents) belonging to Streptococcus (40% of all valves studied), Sphingomonas (35%), Pseudomonas (35%), Roseateles (25%), Phyllobacterium (25%), and Enterococcus (15%). However, for the most part of mitral valves and the small portion of aortal valves there was hardly the variety of bacteria, just one major genus being present: either Streptococcus, Klebsiella or Enterococcus (see the Figure).
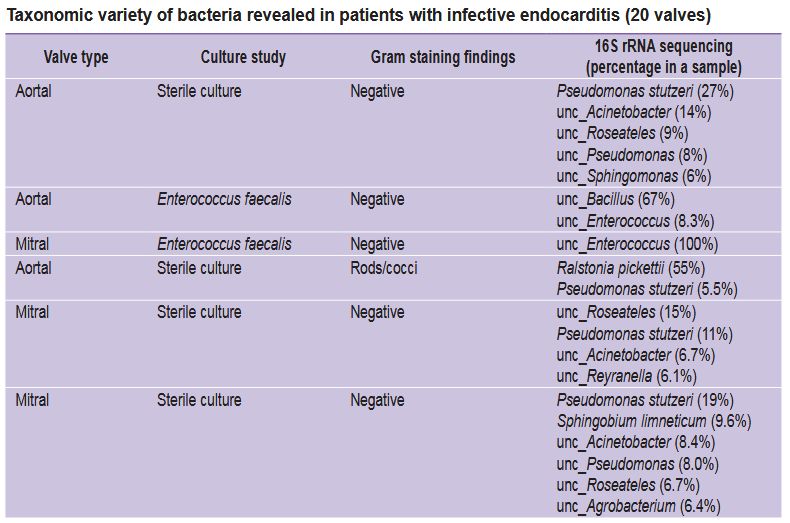
|
Taxonomic variety of bacteria revealed in patients with infective endocarditis (20 valves) |
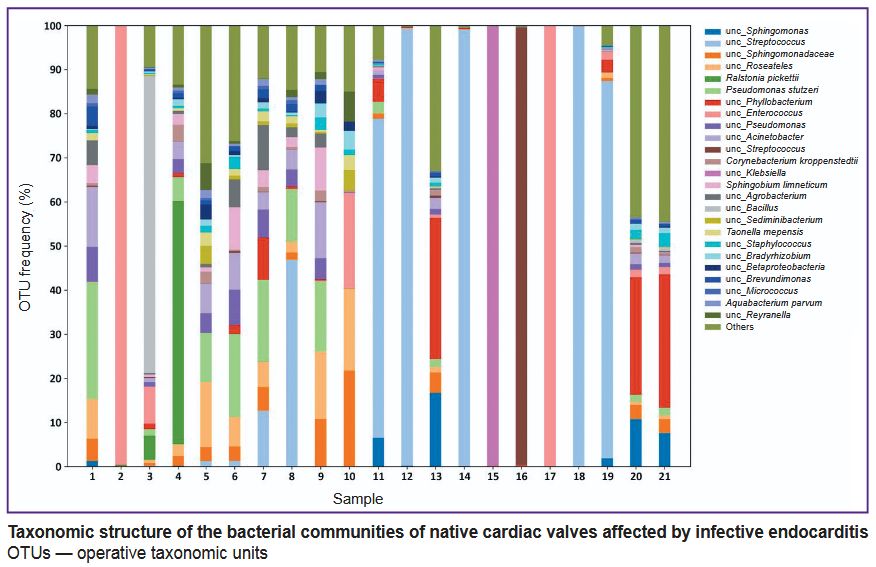
|
Taxonomic structure of the bacterial communities of native cardiac valves affected by infective endocarditis ОТUs — operative taxonomic units |
Discussion
IE is characterized by the microorganisms, which colonize cardiac valves. Despite the advances in the disease prevention, diagnosis and therapy, currently, the fatality rate remains high: nosocomial fatality is from 6 to 50%, 5-year death rate — from 19 to 82% [20]. Epidemiological studies show IE incidence to be from 5.0 to 14.3 cases per 100,000 thousand of adult population [21]. Moreover, IE etiological structure has been found to have changed. So, at present, the main agents (Streptococcus spp. and Staphylococcus spp.) account for nearly 80% of IE cases. In approximately 10% cases the etiological agents are enterococci, particularly, Enterococcus faecalis, and their percentage is higher in patients aged over 65 [22]. Gram-negative rods including those from HACEK group (Haemophilus, Aggregatibacter, Cardiobacterium, Eikenella, and Kingella) are rarely detected (their portion is about 5% of IE cases).
Correct agent identification is a crucially important aspect of IE diagnosis [4]. A conventional definition of IE etiological agent by blood culture or incised tissue can give false-negative results in the range from 2.5 to 31.0% cases; and it is frequently associated with antibiotic therapy prior to admission [20]. In addition, a number of bacteria causing IE can refer to difficult-to-culture or non-cultured groups [23]. The present study demonstrated that the agents revealed by microbiological tests accounted for about 30% of the studied IE cases, while the identification of bacterial agents using 16S rRNA metabarcoding was 100%. So, 16S rRNA enabled to reveal the following bacteria: Streptococcus, Enterococcus, Sphingomonas, Roseateles, Phyllobacterium, Pseudomonas, Ralstonia, Bacillus, and Klebsiella. None Staphylococcus cases were found, despite the fact that these microorganisms are one of the most common agents affecting cardiac valval tissues.
Streptococcus representatives rank next to Staphylococcus aureus in IE pathogenesis. Streptococcus is a heterogeneous group of bacteria containing 50 identified species. Some researchers demonstrate that particular streptococci, primarily from Mitis group, are the main IE agents [24, 25]. Streptococcus gordonii and Streptococcus mutans were shown to be the most common streptococci (they were found in 50% cases), Streptococcus pneumoniae and Streptococcus pyogenes were rarely found (1–2%) [25].
Enterococcus includes gram-positive bacteria holding leading positions among IE agents. Up to 20% of all IE cases can be due to these bacteria, primarily, Enterococcus faecalis [26]. The therapy of IE caused by Enterococcus encounters certain difficulties, since the pathogen is able to form biofilms and preserve microorganisms on both: native cardiac valves, and prosthetic valval structures [27, 28]. It is worth noting that compared to other etiologies, IE caused by E. faecalis is characterized by recurrences after cessation of antibiotic therapy [29, 30].
Moreover, our study revealed two cases of bacterial invasion by rarer microorganisms, such as Ralstonia pickettii and Bacillus spp. R. pickettii belonging to Ralstonia is an aerobic gram-negative oxidase-positive bacillus, which is recently considered as an opportunistic pathogen in patients with compromised immunity. R pickettii can survive in a wide temperature range (from 15 to 42°C), as well as it forms a biofilm resistant to a host immune response and some antibiotics [31]. In addition, currently, there is no standardized protocol to treat infections caused by R. pickettii due to the differences in the sensitivity to antibiotics. Immunosuppressed patients, especially those with acquired (HIV) or pharmaceutical immunosuppression are at risk of getting infected with this type of bacteria [32] that is described by our case of detecting R. pickettii in a patient with involved aortal valve and accompanying HIV and hepatitis C.
In one case 16S rRNA metabarcoding resulted in revealing and identifying Bacillus agent in the patient’s mitral valve, its blood culture showed positive E. faecalis. Bacillus bacteria are gram-positive facultative anaerobes occurring in the environment, food, and in the intestinal microflora [33]. Moreover, Bacillus bacteria have been found to be able to cause IE involving in most cases the cardiac mitral valve, and be associated with heavy mortality in the patients with prosthetic valval structures [34]. Endocarditis caused by Bacillus has been shown to occur frequently in patients with intravenous drug taking, venous catheters, cardiac valval prostheses, malignancies, and immunosuppression [35].
It is of importance to note that the present study revealed one case of gram-negative Klebsiella.The most common Klebsiella species are Klebsiella pneumoniae and Klebsiella oxytoca. Klebsiella granulomatis, Klebsiella ozaenae,and Klebsiella rhinoscleromatis occur rarer. Among the common species it is K. pneumoniae that frequently causes IE, and is characterized by a severe course, mainly due to the fact that these bacteria are resistant to some antibiotics [36].
Conclusion
Thus, the present study findings showed the native cardiac valves excised in IE patients to be characterized by numerous opportunistic pathogenic microorganisms, although blood cultures were negative. It suggests that the residual bacteriemia in patients with preoperative antibacterial therapy aimed primarily at common IE agents can result in developing postoperative prosthetic IE followed by a re-operation, life quality degradation and an increased risk of mortality. The obtained results demonstrated the significance of 16S rRNA metabarcoding of the removed cardiac valves and the necessity for advanced study of taxonomic variety of cardiac valves affected by IE using high-efficiency methods.
Study funding. The study was supported by Russian Science Foundation grant No.23-75-10020 “Molecular and genetic basics of infective endocarditis pathogenesis of native cardiac valves and their bioprostheses”, https://rscf.ru/project/23-75-10020/.
Conflict of interest. The authors declare neither evident nor potential conflict of interest related to the present study.
ОТUs — operative taxonomic units
References
- Li M., Kim J.B., Sastry B.K.S., Chen M. Infective endocarditis. Lancet 2024; 404(10450): 377–392, https://doi.org/10.1016/S0140-6736(24)01098-5.
- Martínez-Sellés M., Muñoz P. Epidemiology, diagnosis, treatment, and prognosis of infective endocarditis. J Clin Med 2023; 12(17): 5705, https://doi.org/10.3390/jcm12175705.
- Stasev A.N., Rutkovskaya N.V., Kokorin S.G., Levadin Yu.V. Salmonella endocarditis of mitral valve: clinical observation. Complex Issues of Cardiovascular Diseases 2017; 2: 123–126, https://doi.org/10.17802/2306-1278-2017-2-123-126.
- Oberbach A., Schlichting N., Feder S., Lehmann S., Kullnick Y., Buschmann T., Blumert C., Horn F., Neuhaus J., Neujahr R., Bagaev E., Hagl C., Pichlmaier M., Rodloff A.C., Gräber S., Kirsch K., Sandri M., Kumbhari V., Behzadi A., Behzadi A., Correia J.C., Mohr F.W., Friedrich M. New insights into valve-related intramural and intracellular bacterial diversity in infective endocarditis. PLoS One 2017; 12(4): e0175569, https://doi.org/10.1371/journal.pone.0175569.
- Demin A.A., Kobalava Zh.D., Skopin I.I., Tyurin P.V., Boytsov S.A., Golukhova E.Z., Gordeev M.L., Gudymovich V.D., Demchenko E.A., Drobysheva V.P., Domonova E.A., Drapkina O.M., Zagorodnikova K.A., Irtyuga O.B., Kakhktsyan P.S., Kozlov R.S., Kotova E.O., Medvedev A.P., Muratov R.M., Nikolaevsky E.N., Pisaryuk A.S., Ponomareva E.Yu., Popov D.A., Rakhina S.A., Revishvili A.G., Reznik I.I., Ryzhkova D.S., Safarova A.F., Tazina S.Ya., Chipigina N.S., Shipulina O.Yu., Shlyakhto E.S., Schneider Yu.A., Shostak N.A. Infectious endocarditis and infection of intracardiac devices in adults. Clinical guidelines 2021. Russian Journal of Cardiology 2022; 27(10): 5233, https://doi.org/10.15829/1560-4071-2022-5233.
- Kotova E.O., Domonova E.A., Kobalava Z.D., Moiseeva A.Y., Pisaryuk A.S., Silveystrova O.Y., Karaulova J.L., Akimkin V.G. Clinical and diagnostic value of including PCR blood test in the traditional algorithm for identifying causative agents of infective endocarditis: a cohort study of 124 patients. Terapevticheskii arkhiv 2023; 95(1): 23–31, https://doi.org/10.26442/00403660.2023.01.202042.
- Zeng X., Wu J., Li X., Xiong W., Tang L., Li X., Zhuang J., Yu R., Chen J., Jian X., Lei L. Application of metagenomic next-generation sequencing in the etiological diagnosis of infective endocarditis during the perioperative period of cardiac surgery: a prospective cohort study. Front Cardiovasc Med 2022; 9: 811492, https://doi.org/10.3389/fcvm.2022.811492.
- Haddad S.F., DeSimone D.C., Chesdachai S., Gerberi D.J., Baddour L.M. Utility of metagenomic next-generation sequencing in infective endocarditis: a systematic review. Antibiotics (Basel) 2022; 11(12): 1798, https://doi.org/10.3390/antibiotics11121798.
- Burban A., Słupik D., Reda A., Szczerba E., Grabowski M., Kołodzińska A. Novel diagnostic methods for infective endocarditis. Int J Mol Sci 2024; 25(2): 1245, https://doi.org/10.3390/ijms25021245.
- Martinez-Porchas M., Villalpando-Canchola E., Ortiz Suarez L.E., Vargas-Albores F. How conserved are the conserved 16S-rRNA regions? PeerJ 2017; 5: e3036, https://doi.org/10.7717/peerj.3036.
- Anton-Vazquez V., Dworakowski R., Cannata A., Amin-Youssef G., Gunning M., Papachristidis A., MacCarthy P., Baghai M., Deshpande R., Khan H., Byrne J., Fife A. 16S rDNA PCR for the aetiological diagnosis of culture-negative infective endocarditis. Infection. 2022;50(1):243-249. doi: https://doi.org/10.1007/s15010-021-01690-x.
- Fadrosh D.W., Ma B., Gajer P., Sengamalay N., Ott S., Brotman R.M., Ravel J. An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2014; 2(1): 6, https://doi.org/10.1186/2049-2618-2-6.
- Shen W., Le S., Li Y., Hu F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS One 2016; 11(10): e0163962, https://doi.org/10.1371/journal.pone.0163962.
- https://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- Edgar R.C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 2013; 10(10): 996–998, https://doi.org/10.1038/nmeth.2604.
- Edgar R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010; 26(19): 2460–2461, https://doi.org/10.1093/bioinformatics/btq461.
- Edgar R.C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 2016, https://doi.org/10.1101/081257.
- Edgar R.C. SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv 2016, https://doi.org/10.1101/074161.
- Wang Q., Garrity G.M., Tiedje J.M., Cole J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 2007; 73(16): 5261–5267, https://doi.org/10.1128/AEM.00062-07.
- Kouijzer J.J.P., Noordermeer D.J., van Leeuwen W.J., Verkaik N.J., Lattwein K.R. Native valve, prosthetic valve, and cardiac device-related infective endocarditis: a review and update on current innovative diagnostic and therapeutic strategies. Front Cell Dev Biol 2022; 10: 995508, https://doi.org/10.3389/fcell.2022.995508.
- Thornhill M.H., Dayer M.J., Nicholl J., Prendergast B.D., Lockhart P.B., Baddour L.M. An alarming rise in incidence of infective endocarditis in England since 2009: why? Lancet 2020; 395(10233): 1325–1327, https://doi.org/10.1016/S0140-6736(20)30530-4.
- Kobalava Z.D., Kotova E.O. Global and national trends in the evolution of infective endocarditis. Kardiologiia 2023; 63(1): 3–11, https://doi.org/10.18087/cardio.2023.1.n2307.
- Reisinger M., Kachel M., George I. Emerging and re-emerging pathogens in valvular infective endocarditis: a review. Pathogens 2024; 13(7): 543, https://doi.org/10.3390/pathogens13070543.
- Cai S., Yang Y., Pan J., Miao Q., Jin W., Ma Y., Zhou C., Gao X., Wang C., Hu B. The clinical value of valve metagenomic next-generation sequencing when applied to the etiological diagnosis of infective endocarditis. Ann Transl Med 2021; 9(19): 1490, https://doi.org/10.21037/atm-21-2488.
- Kim S.L., Gordon S.M., Shrestha N.K. Distribution of streptococcal groups causing infective endocarditis: a descriptive study. Diagn Microbiol Infect Dis 2018; 91(3): 269–272, https://doi.org/10.1016/j.diagmicrobio.2018.02.015.
- Chamat-Hedemand S., Dahl A., Østergaard L., Arpi M., Fosbøl E., Boel J., Oestergaard L.B., Lauridsen T.K., Gislason G., Torp-Pedersen C., Bruun N.E. Prevalence of infective endocarditis in streptococcal bloodstream infections is dependent on streptococcal species. Circulation 2020; 142(8): 720–730, https://doi.org/10.1161/CIRCULATIONAHA.120.046723.
- Habib G., Lancellotti P., Erba P.A., Sadeghpour A., Meshaal M., Sambola A., Furnaz S., Citro R., Ternacle J, Donal E., Cosyns B., Popescu B., Iung B., Prendergast B., Laroche C., Tornos P., Pazdernik M., Maggioni A., Gale C.P.; EURO-ENDO Investigators. The ESC-EORP EURO-ENDO (European Infective Endocarditis) registry. Eur Heart J Qual Care Clin Outcomes 2019; 5(3): 202–207, https://doi.org/10.1093/ehjqcco/qcz018. Erratum in: Eur Heart J Qual Care Clin Outcomes 2020; 6(1): 91, https://doi.org/10.1093/ehjqcco/qcz060.
- Conwell M., Dooley J.S.G., Naughton P.J. Enterococcal biofilm — a nidus for antibiotic resistance transfer? J Appl Microbiol 2022; 132(5): 3444–3460, https://doi.org/10.1111/jam.15441.
- Lecomte R., Laine J.B., Issa N., Revest M., Gaborit B., Le Turnier P., Deschanvres C., Benezit F., Asseray N., Le Tourneau T., Pattier S., Al Habash O., Raffi F., Boutoille D., Camou F. Long-term outcome of patients with nonoperated prosthetic valve infective endocarditis: is relapse the main issue? Clin Infect Dis 2020; 71(5): 1316–1319, https://doi.org/10.1093/cid/ciz1177.
- Calderón-Parra J., Kestler M., Ramos-Martínez A., Bouza E., Valerio M., de Alarcón A., Luque R., Goenaga M.Á., Echeverría T., Fariñas M.C., Pericàs J.M., Ojeda-Burgos G., Fernández-Cruz A., Plata A., Vinuesa D., Muñoz P.; on behalf of the GAMES investigators. Clinical factors associated with reinfection versus relapse in infective endocarditis: prospective cohort study. J Clin Med 2021; 10(4): 748, https://doi.org/10.3390/jcm10040748.
- Danneels P., Hamel J.F., Picard L., Rezig S., Martinet P., Lorleac’h A., Talarmin J.P., Buzelé R., Guimard T., Le Moal G., Brochard-Libois J., Beaudron A., Letheulle J., Codde C., Chenouard R., Boutoille D., Lemaignen A., Bernard L., Cattoir V., Dubée V; EFEMER study group. Impact of Enterococcus faecalis endocarditis treatment on risk of relapse. Clin Infect Dis 2023; 76(2): 281–290, https://doi.org/10.1093/cid/ciac777.
- Basso M., Venditti C., Raponi G., Navazio A.S., Alessandri F., Giombini E., Nisii C., Di Caro A., Venditti M. A case of persistent bacteraemia by Ralstonia mannitolilytica and Ralstonia pickettii in an intensive care unit. Infect Drug Resist 2019; 12: 2391–2395, https://doi.org/10.2147/IDR.S206492.
- Orme J., Rivera-Bonilla T., Loli A., Blattman N.N. Native valve endocarditis due to Ralstonia pickettii: a case report and literature review. Case Rep Infect Dis 2015; 2015: 324675, https://doi.org/10.1155/2015/324675.
- Wright W.F. Central venous access device-related bacillus cereus endocarditis: a case report and review of the literature. Clin Med Res 2016; 14(2): 109–115, https://doi.org/10.3121/cmr.2016.1312.
- Gopinathan A., Kumar A., Sen A.C., Sudha S., Varma P., Gs S., Eapen M., Dinesh K.R. A case series and review of bacillus cereus endocarditis from India. Open Microbiol J 2018; 12: 28–33, https://doi.org/10.2174/1874285801812010028.
- Ioannou P., Miliara E., Baliou S., Kofteridis D.P. Infective endocarditis by Klebsiella species: a systematic review. J Chemother 2021; 33(6): 365–374, https://doi.org/10.1080/1120009X.2021.1888025.










